AMR Genomics Research Group
School of Medical, Molecular and Forensic Sciences
Murdoch University, Australia
Group Leader
Antimicrobial resistance (AMR) is a growing global health threat.
The AMR Genomics Research Group investigates the genomic epidemiology and mechanisms of AMR in microbial pathogens using whole-genome sequencing, advanced bioinformatics, and molecular approaches. Through this integrated framework, we uncover how resistance emerges and spreads - bridging microbiology, genomics, and public health.

Recent Research Highlights
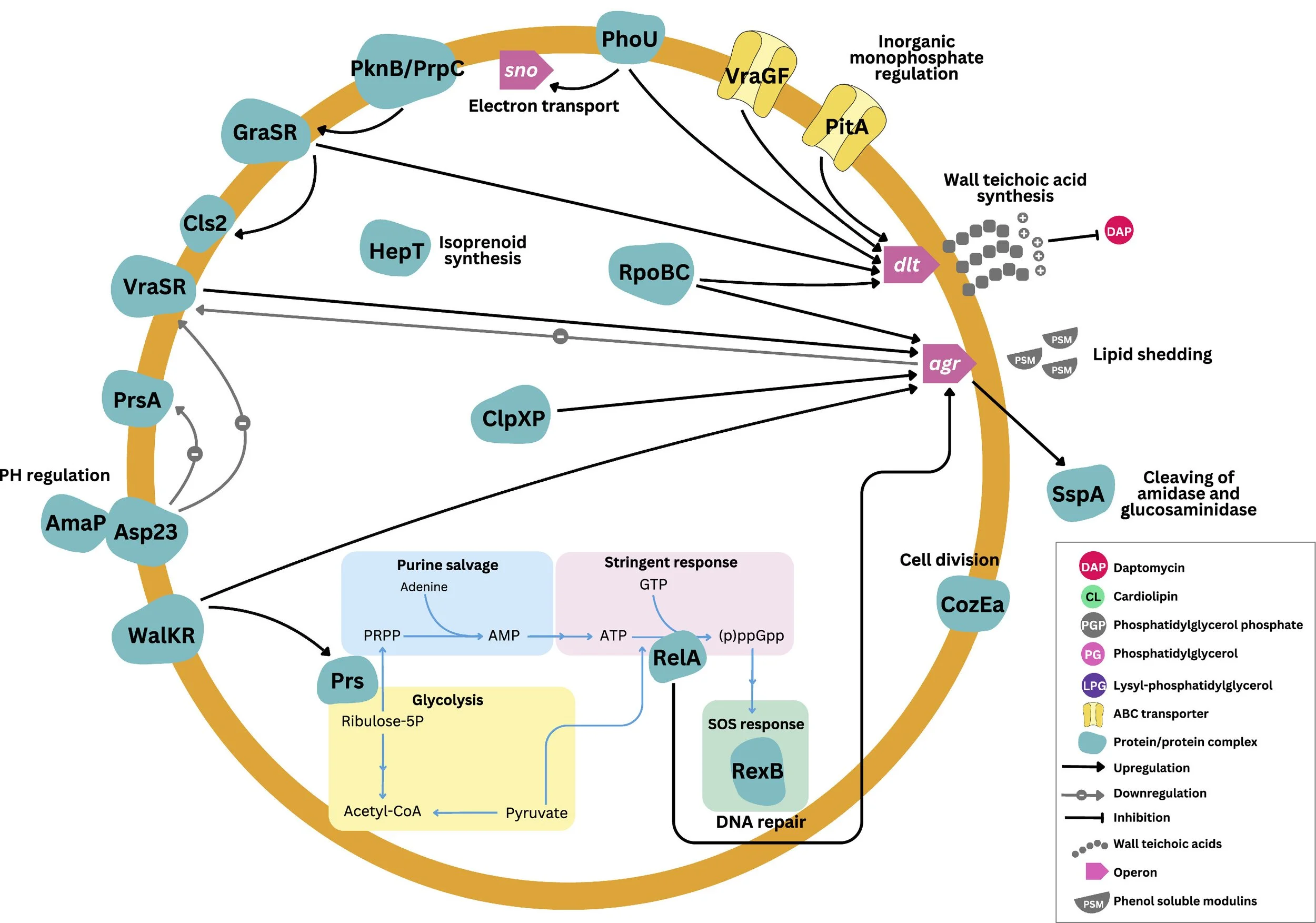
Molecular Mechanisms of Daptomycin Resistance in S. aureus
Daptomycin is a last-resort antibiotic used to treat infections caused by multidrug-resistant Gram-positive pathogens, including methicillin-resistant Staphylococcus aureus (MRSA). However, reports of daptomycin resistance in S. aureus are increasing globally. Resistance is commonly associated with alterations in membrane lipid composition, particularly through pathways regulating lysyl-phosphatidylglycerol synthesis controlled by two-component systems. In addition, peptidoglycan modifications may disrupt penicillin-binding protein activity, contributing to the daptomycin–β-lactam “see-saw” effect. Importantly, daptomycin tolerance—often observed in small colony variants and biofilms—can act as a precursor to the development of stable resistance.
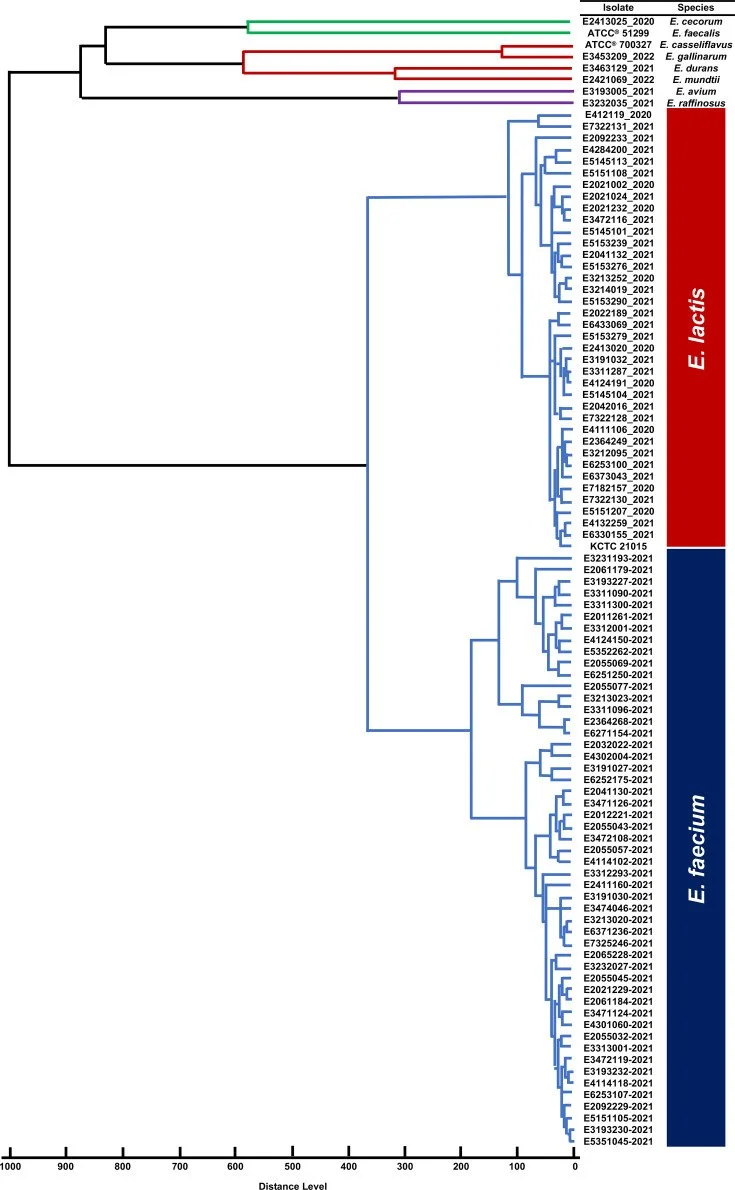
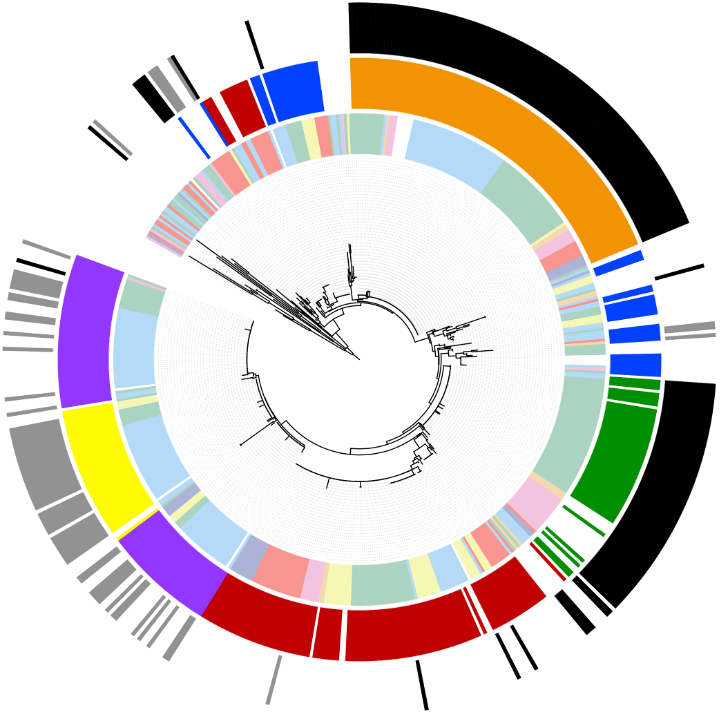
Accurate identification of Enterococcus lactis causing bacteraemia by matrix-assisted laser desorption ionization-time of flight mass spectrometry
Following the reclassification of Enterococcus faecium clade B as Enterococcus lactis, accurate identification of this species has become increasingly important, particularly in clinical settings where it may be misidentified. In this study, a custom MALDI Biotyper® database was developed and validated using whole-genome sequence-confirmed isolates to enable rapid differentiation of E. lactis from E. faecium. The in-house database achieved 100% correct species-level identification, successfully distinguishing E. lactis isolates that were otherwise misidentified by standard commercial databases. This approach provides a practical and reliable tool for improving diagnostic accuracy in clinical and research laboratories.